Engineering
Introduction to DBTL
Engineering is a set of principles that we use to effectively design and build projects of interest. When developing a biological system, this framework consists of four stages, commonly referred to as the "Design-Build-Test-Learn cycle" (DBTL cycle) . We embrace this cycle as an engineering principle to address real-world problems or achieve specific goals. The following are key stages within the DBTL cycle:
DBTL cycle is an iterative process that provides opportunities for understanding and improving biological systems. It serves as a fundamental engineering principle that runs through our project “Star-Chlamy” and plays a crucial role. In response to the urgent need to address the agricultural crisis and achieve sustainable development goals, we have designed an innovative approach using microalgae as cell factories for starch production.
To achieve our proof-of-concept, we have gone through multiple DBTL cycles that encompass various aspects of our project design. Through continuous design, building, testing, and learning, we have progressively improved and optimized the functionality and performance of the selected biological system, enabling Star-Chlamy to better serve real-world demands and applications.
Figure 1.2 An overview of our project “Star-Chlamy”
Design
Chassis Selection
Chassis cells serve as the "hardware" foundation of synthetic biology. Based on our objectives, we have chosen Chlamydomonas reinhardtii as the chassis for our project. Chlamydomonas reinhardtii is a unicellular eukaryotic photosynthetic microalga that offers the following advantages:
Pathway selection
We have employed the principles of metabolic engineering in our design. The cellular factory aims to achieve optimized synthesis of the entire metabolic pathway by modulating the native starch metabolism of the cell, effectively allocating light and carbon outputs.
Scientists have elucidated the starch metabolism pathway in Chlamydomonas reinhardtii (Radakovits et al., Eukaryotic Cell, 2010), which has provided us with considerable convenience. Based on this, we have decided to target this pathway for our project.
Additionally, considering the crosstalk between the starch metabolism pathway and the lipid metabolism pathway, we have also targeted the lipid pathway to indirectly promote starch synthesis.
Wet Lab Design
After selecting the pathway, we proceeded to design our laboratory procedures. We devised a strategy utilizing the principle of "broaden the source of income and economize on expenditure" to simultaneously promote starch synthesis and inhibit starch degradation.
The Star-Chlamy project is a classic example of metabolic engineering. In metabolic engineering, certain pathways are often limited at two or more metabolic nodes. A feasible solution to this issue is to combine regulatory modules to control the metabolic flux at multiple nodes. Therefore, we have divided the starch metabolic pathway into three major modules, following an engineering approach: modifying starch metabolic pathway(Regulatory module 1.1-1.4)、expand the optical absorption spectrum (Regulatory module 2)、broaden the application scenarios of CCM (Regulatory module 3). By combining these three modules, we can control the output of targeted metabolic circuit.
Regulatory module 1.1:
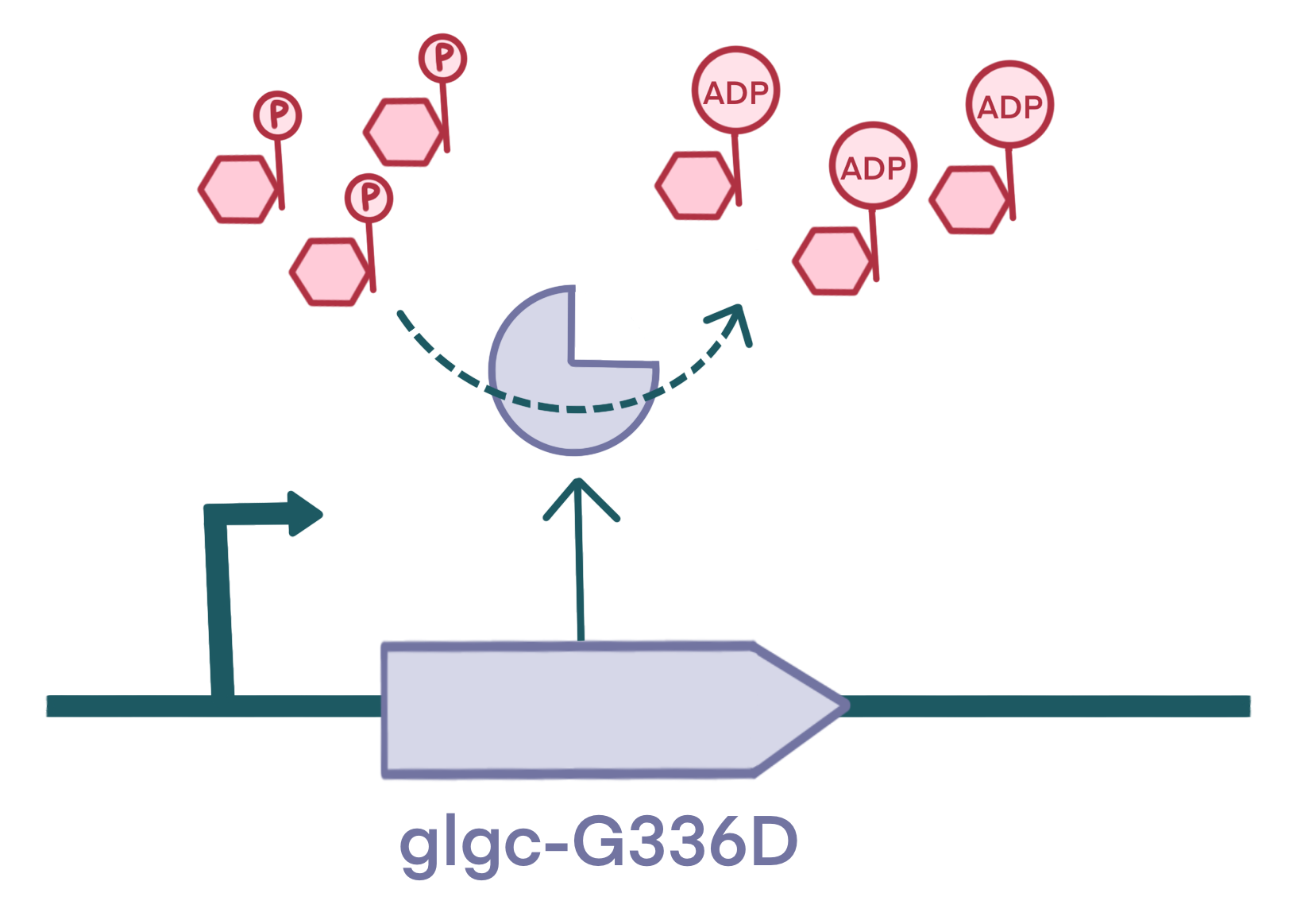
Metabolic engineering often alters production by altering enzyme levels. Through literature research, we discovered that AGPase is the rate-limiting enzyme instarch synthesis. So we designed to introduce a more efficient AGPase encoded by glgC from Escherichia coli to AGP4-knockout chlamy strain. In this way, the source of income can be broaden.
Regulatory module 1.2:
It is also important to economize on expenditure in engineering thinking. GWD plays a crucial role in the process of starch degradation. Consequently, we selected the corresponding genes GWD1 and GWD2 as targets and manipulated them accordingly.
Regulatory module 1.3:
At the same time, reducing the competitive endogenous pathways can also improve the production. Since starch synthesis and lipid synthesis compete with each other, we decided to inhibit lipid synthesis to promote starch synthesis. Due to the inherent uncertainty of this approach, we have chosen not to manipulate genes at this stage. Instead, we are designing a cultivation experiment using the lipid synthesis inhibitor, cerulenin, on the chassis Chlamy.
Regulatory module 1.4:
Regulation of the value of metabolic products is also important. After communicating with the China Starch Industry Association(Click Here to see more details), we learned that high amylose starch has higher added value. To promote the accumulation of high amylose starch, we decided to target the SBE enzyme and knock out the SBE2 and SBE3 genes.
Regulatory module 2:
Starch is one of the important storage carbon sources, and it is crucial to improve the yield of starch from the source. Therefore, we have chosen the key node, right reaction, as module 2. In runway ponds for industrial production and cultivation of Chlamydomonas, Chlamy in the upper layer uses near-red light for photosynthesis. Therefore, a serious problem of near-red light attenuation in the depth of runway ponds exists, which leads to serious decrease of the photosynthetic efficiency of Chlamydomonas in the depth of runway ponds. So we targeted it to broaden the light absorption spectrum.
Regulatory module 3:
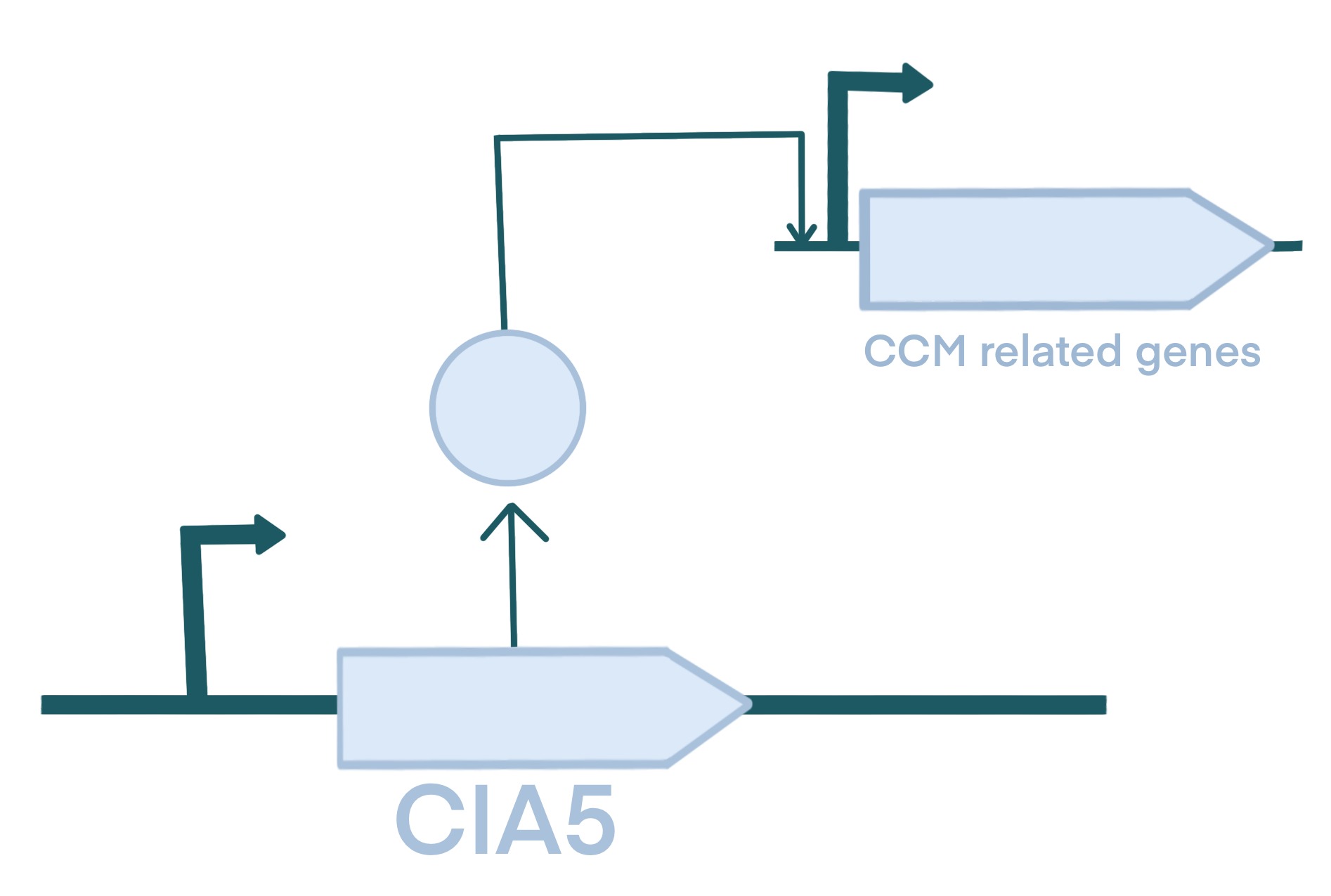
In addition to that, we also target the dark reactions of chlamy photosynthesis. Our chassis, Chlamydomonas reinhardtii, possesses a carbon concentration mechanism (CCM) that allows for the high concentration enrichment of CO2 even in low external inorganic carbon conditions, giving it a certain advantage. However, this mechanism is inhibited under high CO2 concentrations. Therefore, in order to further increase starch metabolism flux, we designed to regulate the key transcription factor CIA5 of CCM.
In Silico Design
In addition, we use computer simulation and mathematical modeling to help optimize the design and also verify if the laboratory design functions correctly.
Module 1 - starch pathway
First, we modeled starch metabolism based on the Mie equation to assess how the activity our targets affects the amount of starch accumulated. We focused on the two main variables that make changes in the amount of starch accumulated, GWD as well as AGPase activity. Using the Matlab, we can get the results, which matches with the expective results in the wet lab design.
We also model the starch branching process for verification. Differential equations are used to represent the starch branching kinetics and give the starch branching profile. From the results it can be seen that the larger the relative activity of SBE, the smaller the amount of straight chain starch. The wet lab designs are evaluated and verified through data analysis.
Module 3-CCM
In order to provide ideas for our operation of CIA5, we start with modeling CIA5's 3D structure. It has been reported that the CIA5 protein gene with a deletion of 54 amino acids in the C terminus can complement the CIA5 mutant phenotype[2], which suggests that the C terminus of CIA5 may function as the upstream CO2 sensing domain and regulate its own transcription factor activity. There is currently a lack of CIA5 structure information on the Internet, so we used Alpha Fold to predict CIA5 and C terminus deleted CIA5 structures.
The prediction showed that: The C terminus was located on the lateral side of the protein. The confidence of the C-terminal structure prediction was less than 50%, and it was speculated to be a flexible structure (it could not be excluded that the lack of homologous protein structure information makes it difficult to predict, resulting in low confidence). And truncated CIA5 center relatively stable structure before and after almost no change. The external flexible part also changed less. So this suggests that the C-terminal itself may not be an influencing factor, but the C-terminal phosphorylation reported by predecessors plays a role.
Therefore, we analyzed the C end of potential phosphorylation sites of CIA5 using NetPhos 3.1[3] . Those with high scores were highly likely. Among the 54 amino acids in the C-terminal region of CIA5, there were three potential phosphorylation sites, 649T, 667S and 696S, and the highest scoring phosphorylase were cdc2, PKA and CKⅡ, respectively. The highest rating of the three was 667S. This provides ideas for our subsequent experimental operations in the Build stage.
Build
DNA Synthesis
During Build stage, the designs are implemented through specific experiments to construct the required biological systems. Based on the targets identified in the Design stage, we constructed several basic parts.
Module 1-starch pathway
For regulation module 1.1, we built part BBa_K4772000:glgC-G336D . Its gene sequence was derived from a more efficient modified AGPase (referred as glgC-G336D) from Escherichia coli, which was optimized using preferred codons for Chlamydomonas reinhardtii. Additionally, to ensure standardization of our basic part, we avoided using restricted restriction enzyme recognition sites. We synthsized the BBa_K4772000: glgC-G336D DNA by company and performed PCR for amplification.
Module 2-light absorption
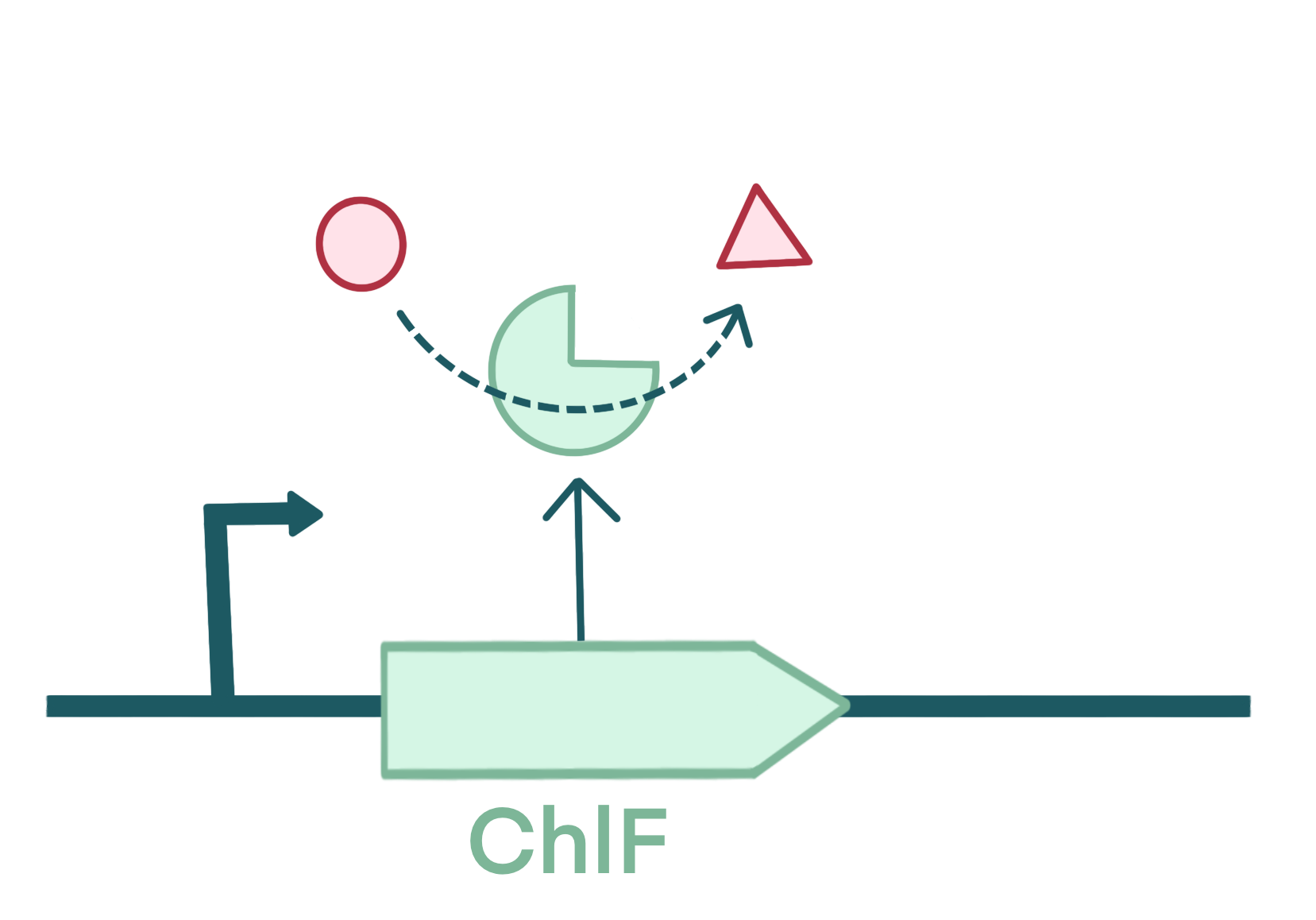
For module 2, we built part BBa_K4772001: ChlF. It encodes enzyme ChlF from cyanobacteria, which produces chlorophyll f (Chl f) by oxidizing chlorophyll a (Chl a). Chlamydomonas reinhardtii are organisms that convert visible photons into chemical energy via oxygenic photosynthesis. And it usually uses Chl a as photochemically active pigment. However, Chl f, as a red-shifted pigment, has been discovered recently[4]. It was found in terrestrial cyanobacteria and for evolution pressure, they have evolved a novel far-red light photoacclimation (FaRLiP) use far-red light (FR) for photosynthesis. So express ChlF in Chlamydomonas may help to extend their light harvesting and improve their photosynthetic light use efficiency. We obtained the BBa_K4772001: ChlFChlF DNA through commercial synthesis and performed PCR for amplification.
Module 3-CCM
For module 2, we built part BBa_K4772002:CIA5 . It encodes an artificial CIA5 mutant with a C-terminal truncation into the wild type Chlamydomonas reinhardtii to allow it to function as a activated transcription factor under any CO2 concentration. Since CIA5 gene segment is relatively long, our funding did not support its commercial synthesis. Also, because of the high GC content of Chlamydomonas reinhardtii, we could not directly amplify CIA5 from its genome. Therefore, we divided CIA5 gene into three segments and amplified them separately, which were then spliced by overlapping PCR. Finally, We obtained the BBa_K4772002: CIA5 DNA.
Part Assembly
To assemble the BioBricks,we constructed several plasmids to construct genetic circuits. The YFP tag made sure that the expression of basic parts can be verified by performing Western Blot. PsaD CTP is a chloroplast transport peptide that helps transport the parts into chloroplasts to function. And the HA tag made sure that the expression of parts can be verified by performing Western Blot.
We used the seamless assembly closing kit from company CloneSmarter to assemble the synthesized parts onto the plasmid vector using homologous recombination. By employing this method, we successfully constructed these plasmids and obtained accurate sequencing results.
Gene Editing
In the upcoming chassis modification, although genetic editing of Chlamydomonas has been proven challenging worldwide, we have successfully overcome this obstacle by utilizing a gene editing system developed by the collaborative laboratory of professor JUNMIN PAN. We employed CRISPR/Cas9technology to knockout the AGP4, GWD1, GWD2, SBE2, and SBE3 genes in both the 21gr and srt2 Chlamydomonas cell lines (considered wild-type). After obtaining individual gene knockout mutants, we further constructed double knockout mutants, achieving the presence of both mutations within the same algal strain. Besides, we introduced glgC-G336D、ChIF and CIA5 plasmids into Chlamydomonas using electroporation.
Cell Treatment
For lipid synthesis inhibition, we set a control group and an experimental group. The control group was supplemented with 0.1% methanol, while the experimental group was supplemented with 10 μM cerulenin in 0.1% methanol. After 48 hours of nitrogen-depleted culture, we measured the starch content.
Test
Module 1 Modificating Starch Metabolic Pathway
1) Screening
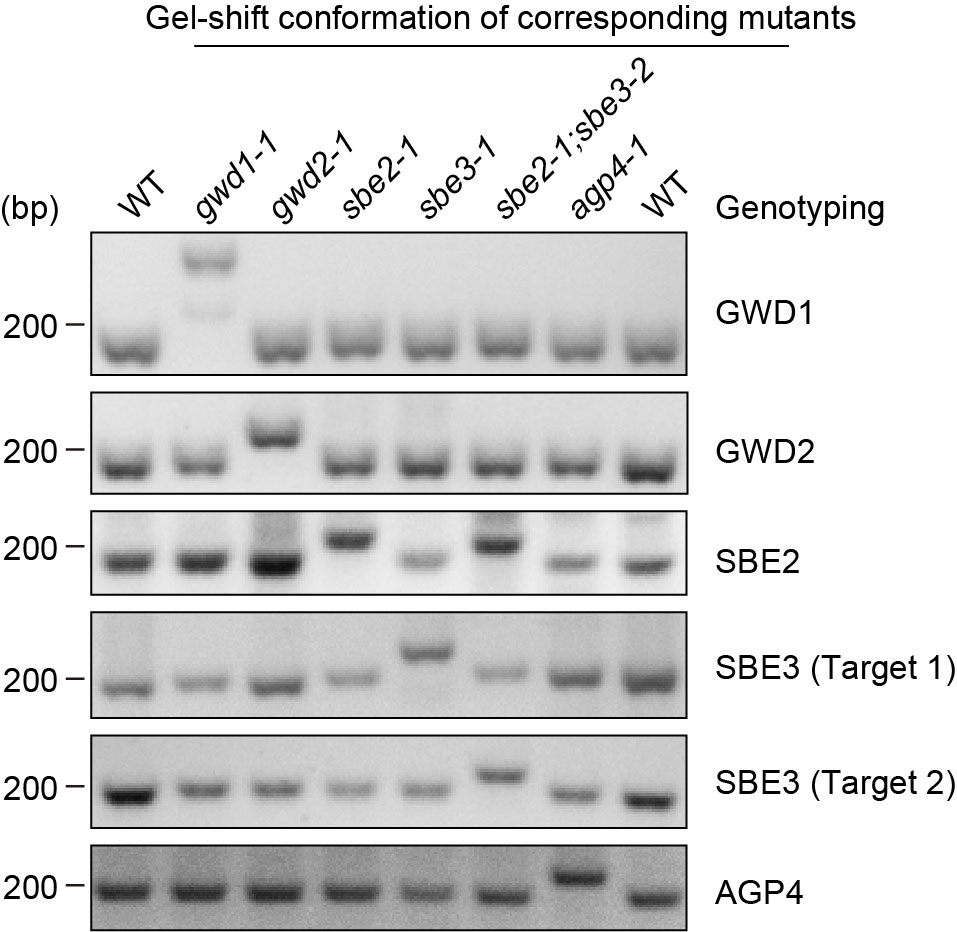
Once the system is built, we experimentally validate the constructed chassis. We performed genotyping and successfully obtained five positive strains respectively for targeted genes. After further check, we confirm that they are knocked out mutant. Additionally, we also screened for a mutant with double knockout of SBE2 and SBE3, thereby achieving the presence of these two mutations in the same strain.
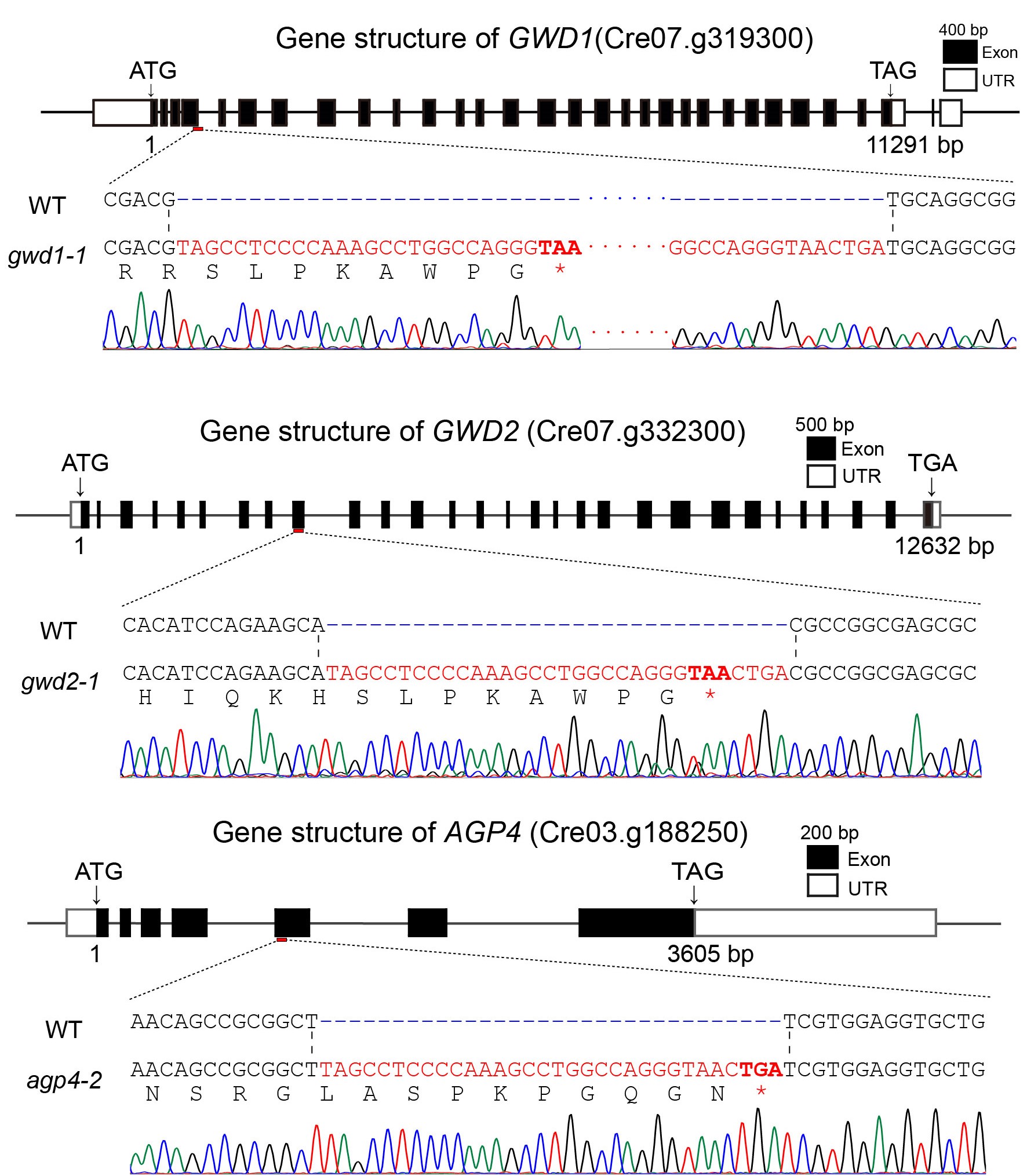
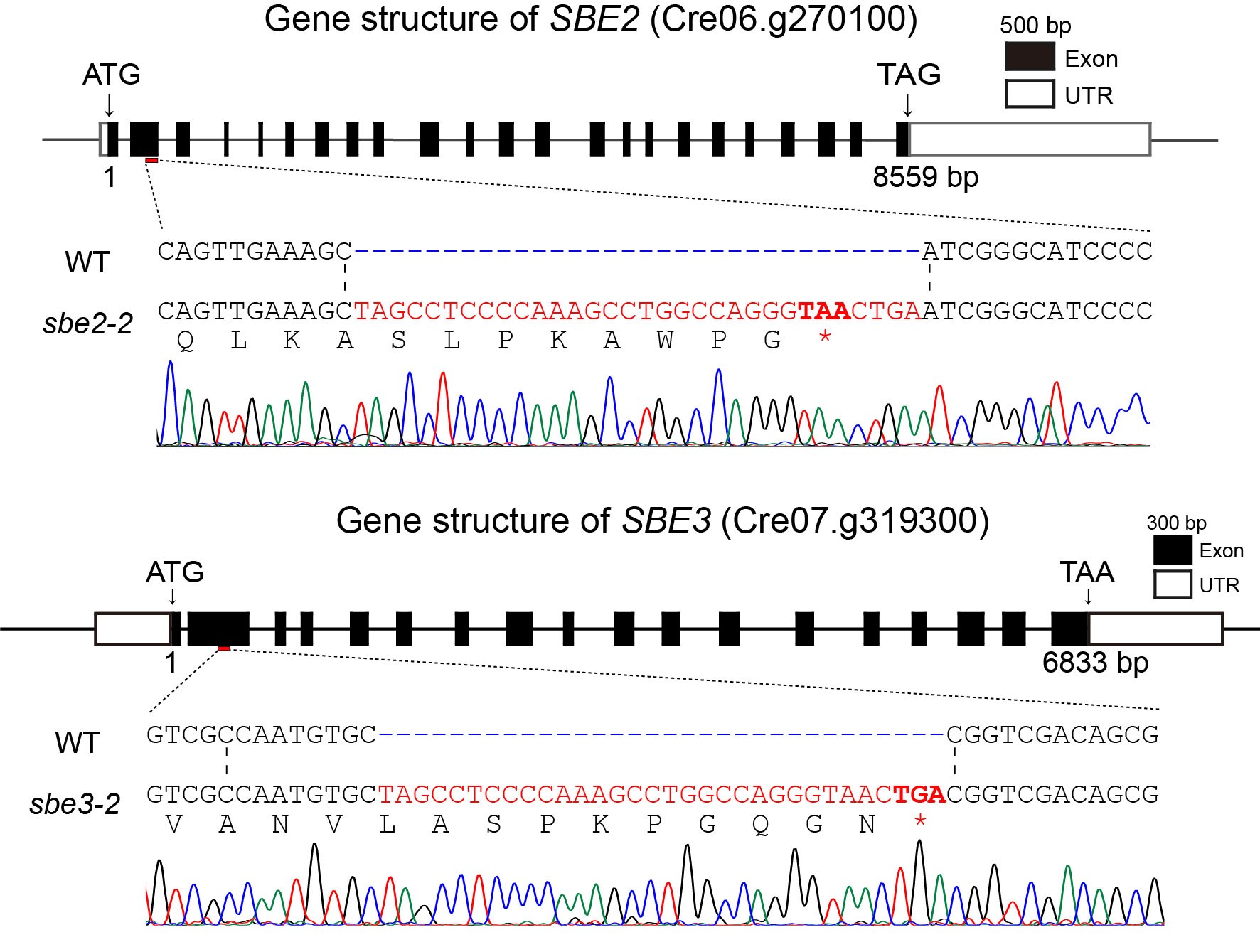
Figure 3.3 Gene insertion and sequencing results of the five mutant strains
In the sequencing results, the red, bolded sequences with asterisks represent the inserted stop codons. All five mutant strains successfully inserted stop codons, resulting in disrupted gene expression and successful gene knockout. Also, we got a SBE double mutant sbe2-3;sbe3-5 by carrying out gene knockout in obtained mutant.
Besides, we conducted fluorescence detection using YFP tags and screened in the 24-well plates. We obtained two positive strains , each transferred one type of glgC-G336D plasmids(PsaD CTP-glgC G336D-mVenus and glgC G336D-YFP).
Figure 3.4 Results of fluorescence intensity detection
Excitation light wavelength: 488nm
Absorption light wavelength range: 500-520nm
The two 24-well plates in the first row show 48 different cell clones introduced into the PsaD CTP-glgC G336D-mVenus vector, and the second row both show 48 different cell clones introduced into the glgC G336D-YFP vector. One positive cell line was obtained from each of the two vectors.
Analytical chemistry
To test the effectiveness of our strategy, we have developed a method for extracting and quantifying starch content in Chlamydomonas based on literature research and constant trial. This method allows us to measure the starch content in both the wild-type strain and the mutants, thereby validating whether there is an increase in starch content. Additionally, we have utilized a commercial assay kit to determine the proportion of linear starch in the mutants, specifically measuring the increase in linear starch content. Furthermore, we referred to literature and captured scanning electron microscope (SEM) images of starch granules to provide a visual representation of the proportion of linear starch within the cells.
By employing these techniques, we aim to obtain quantitative data on starch content and assess the changes in linear starch proportions in the mutants compared to the wild-type strain.
1.Starch content standard curve
The standard curve was drawed using starch sample with a series of concentration gradients in the test
kit(Starch Content Assay Kit (BOXBIO, China, AKSU015C)). The curve would be used in the concentration of
starch produced by the sample we wanted to measure.
The horizontal axis represents the concentration of the standard starch samples and the vertical axis represents their absorbance at △620nm. The data were analyzed by linear regression. The linear regression formula is △620 = 6.614 Concentration. R2 = 0.9994.
2.Total starch content of cells cultivated by lipid synthesis inhibitor
According to the previous research using cerulenin in Chlamydomonas[6], we
designed a control group and an experimental group. The control group was supplemented with 0.1% methanol,
while the experimental group was supplemented with 10 μM cerulenin in 0.1% methanol. After 48 hours of
nitrogen-depleted culture, we measured the starch content.
Figure 3.5 Qualitative and quantitive starch content of control and cerulenin treatment group
It can
be seen from the figure that the starch content of cerulenin treatment group was significantly higher than
the control group, about 2 times that of wild type.
3.Total starch content of six mutants
According to previous research[7], we measured the starch content of mutants
after 5 days of nitrogen-depleted culture. 5×10^7 cells of wild type and each six mutants were collected
for measurement.
Figure 3.6 Qualitative and quantitive starch content of wild type and all mutants
It can be seen from the figure that the starch content of mutant gwd1-1 and gwd2-1 were higher than wild type, but gwd1-1 was significant while gwd2-1 was not. The starch content of mutant sbe2-3 and sbe2-3; sbe3-5 were significantly higher than wild type, in which sbe2-3 was 7.91 times that of wild type. But sbe3-4 was lower than wild type(not significant). agp4 was significantly lower than wild type.
Amylose content and perception
We use the Amylose Content Assay Kit ( Solarbio, China, BC4265) to measure amylose content. The starch sample for measurement were from the starch we extracted from cells before.
1.Amylose content of mutant sbe2-3/sbe3-4/sbe2-3;sbe3-5
Figure 3.7 Qualitative and quantitive amylose content of wild type and SBE mutants
It can be seen from the figure that the amylose content of 3 kinds of mutants were higher than wild type. sbe2-3 and sbe2-3; sbe3-5 were significantly while sbe3-4 was not. sbe2-3 was 8.82 times that of wild type.
2.Amylose perception of mutant sbe2-3/sbe3-4/sbe2-3;sbe3-5
It can be seen from the figure that amylose perception of mutant sbe3-4 and sbe2-3;sbe3-5 were significantly higher than wild type, and sbe2-3;sbe3-5 double mutant’s amylose perception was about twice that of wild type. sbe2-3 was also higher but not significant. The result was consistent with the research paper[8].
Phenotypic characterization of starches produced by mutant
The figures were observed using SEM. It can be seen that in wild type cells, the surface of starch particles were round and smooth like that in the previous article[8]. But in mutant cells the particles were folded and angular, which showed that the phenotypic characterization changed significantly.
3) Cell characteristic
In order to assess whether the gene knockout affects the normal growth of Chlamydomonas, we conducted a three-day growth curve experiment with several obtained mutants, using the wild-type strain as a control. The growth curve was measured at 8-hour intervals using optical density (OD) readings as a classic method for characterizing cell density. OD values of the Chlamydomonas culture were measured at 16:00, 24:00, and 8:00, respectively. The growth curves were plotted, and the differences were calculated.
By comparing the growth curves and analyzing the differences in OD values, we can evaluate the impact of the gene knockout on the growth of Chlamydomonas and determine if there are any significant deviations from the wild-type control.
Module 2 Extend light harvesting of Chlamy
1) Screening
To screen for positive strains for ChIF, we conducted fluorescence detection using YFP tags and screened
in the 24-well plates. We obtained one positive strain transferred the plasmid PsaD CTP-ChIF-mVenus.
Analytical chmistry
By performing Fluorescence intensity detection and Western Blot, we can verify that ChlF was successfully
expressed in our Chlamy.
(Excitation light wavelength: 488nm
Absorption light wavelength range: 500-520nm)
The first and second lines are two repeats. Samples in the same columns but different lines come from one strains of algae. Scale of the heat map is shown on the right. NC: Chlamy that do not express YFP; PC: Chlamy that express YFP; ChlF-YFP: Chlamy that express ChlF-YFP plasmid. ChlF-YFP-1 and ChlF-YFP-2 are two different strains of Chlamy.
Module 3- Broaden the application scenarios of CCM
1) Screening
Module 3 Broaden the application scenarios of CCM
Western blot method was applied to verify the expression of the CIA5ΔC plasmid. Results showed no HA tag expression was observed in all 96 lines examined, which means that screening for the target phenotype should continue. Lane 1-12: Chlamy that fail to expression CIA5ΔC; PC: positive control; 1st Ab: Rat anti-HA; 2nd Ab: Goat anti-rat.
Actually, before we constructed the CIA5-C-del plasmid, we had tried to knock out the CIA5 gene at 4 different target site as well as reproduce the C-terminus truncation mutant by CRISPR method, aiming at confirming its CO2 responding function and transcription factor activity. A total of 1,008 algae strains after CRISPR operating had been tested by genotyping, with no positive result showing a successful editing on CIA5 gene. This suggests that: 1) CIA5 is strongly required for Chlamydomonas life activities and CIA5 knockout may lead to cell death; 2) The chromatin status of CIA5 is not suitable for CIRSPR system for gene editing.
Either way, we have provided a good topic for future iGEM teams to pursue Chlamydomonas related research.
Learn
During Learn stage, an analysis is conducted on the data from Test to acquire principles that facilitate the more effective advancement of the cycle towards desired objectives. These principles serve as guidance for improving the design in the subsequent cycle. To achieve our proof-of-concept, we have gone through multiple DBTL cycles that encompass various aspects of our project design. Through Learn, we progressively optimized the Design, Build and Test , improving performance of the selected biological system and enabling “Star-Chlamy” to better serve real-world demands.
For Design
Learn provides a perspective for our future Design.
In our experimental results, the starch content of sbe2 mutant was significantly higher than that of the wild type, while sbe3 mutant had less starch. Interestingly, during the mid-term of our project, a paper newly published on 31st May 2023 reported the knockout of SBE2 and SBE3 in Chlamydomonas reinhardtii, but their results were not entirely consistent with ours. According to the paper, sbe3 mutant showed a reduction in starch content, while the sbe2 mutant showed little change. In light of this, we speculate that the starch accumulation in our sbe2 mutant, which is a short cilium strain with impaired motility, might be due to the movement defect. This observation has inspired us to consider a possible association between movement defects and starch accumulation. It provides us with a direction for future pathway analysis, considering the incorporation of genes related to movement defects in our design.
Furthermore, our attempts to culture Chlamydomonas with inhibitors of lipid synthesis have shown significant success. This inspires us to consider genetic manipulation of the lipid synthesis module(Regulation module 2) in the future.
Because of time or other reasons, our results showed no HA tag expression was observed in all 96 lines examined, which means that screening for the target phenotype should continue.
However, it also suggests to us that CIA5 might not be easy to express in Chlamydomonas. Therefore, we can learn from this and plan to explore other genetic elements, such as knocking out the downstream binding site of CIA5, to overcome this challenge.
For Build
Based on the results from previous rounds of CIA5, we have developed an interest in the binding site downstream of CIA5. We have constructed an LCR1 gene expression plasmid and inserted it into Chlamydomonas genome, and to screen for a line with constitutively high LCR1 gene expression (relative to wild type). We hope that this could further enhance CCM protein expression in Chlamydomonas on the basis of the CIA5-C-del strain to further enhance its CO2 assimilation capacity. Relevant experiments are still in progress.
Learn not only help us gain insights for Design, but also provides guidance for Build.
Initially, we intended to construct double mutants such as gwd1; sbe3 to synthesize starch that is both abundant and linear. However, to our surprise, we found that the starch content in the sbe2 mutant significantly increased, even surpassing the levels observed in the gwd1 mutant and gwd2 mutant. As a result, we decided not to construct double mutants like gwd1; sbe3 and instead directly constructed the sbe2; sbe3 double mutant. This decision was based on what we learned through practical experimentation.
For Test
Learn can also be beneficial for optimizing Test and generating better screening methods. In the initial design round, we constructed glgC G336D-HA but did not obtain positive results for the HA tag through screening. Learning from this failure, we introduced YFP and utilized fluorescence-assisted screening, which improved the efficiency of our screening process. This approach indeed guided us in obtaining positive strains of glgC-G336D.
Besides, to find a more efficient screening method, we constructed part BBa_K4772003: SGT1 using the GCaMP fragment provided by He Yue from Professor Junmin Pan's Lab. Chlamydomonas mutants of SGT1 have cilia-related defects and thus agglomerate in TAP medium and unable to move. Based on this characteristic, we are able to use SGT1 as a screening marker to screen Chlamydomonas reinhardtii strains that successfully express foreign proteins, which provided us methods for further concept validation.
Reference:
[1]. https://s10.lite.msu.edu/res/msu/botonl/b_online/e19_1/metabol.htm
[2]. Xiang, Y et al. The Cia5 gene controls formation of the carbon concentrating mechanism in Chlamydomonas reinhardtii. National Academy of Sciences of the United States of America vol. 98,9 (2001): 5341-6.
[3]. NetPhos 3.1 - DTU Health Tech - Bioinformatic Services.
[4]. Chen, M. et al. A red-shifted chlorophyll. Science 329, 1318–1320 (2010).
[5]. https://sites.dartmouth.edu/dujs/2021/01/24/beginning-stages-of-crispr-detection-in-athletes/
[6]. Luis Gonzaga Heredia-Martínez, Ascensión Andrés-Garrido, Enrique Martínez-Force, María Esther
Pérez-Pérez, José L. Crespo, Chloroplast Damage Induced by the Inhibition of Fatty Acid Synthesis
Triggers Autophagy in Chlamydomonas, Plant Physiology, Volume 178, Issue 3, November 2018, Pages
1112–1129, https://doi.org/10.1104/pp.18.00630
[7]. Delrue, B., Fontaine, T., Routier, F., Decq, A.,
Wieruszeski, J. M., Van Den Koornhuyse, N., Maddelein, M. L., Fournet, B., & Ball, S. (1992). Waxy
Chlamydomonas reinhardtii: monocellular algal mutants defective in amylose biosynthesis and granule-bound
starch synthase activity accumulate a structurally modified amylopectin. Journal of bacteriology, 174(11),
3612–3620. https://doi.org/10.1128/jb.174.11.3612-3620.1992
[8]. Courseaux, Adeline et al. “BE3 is the major branching enzyme isoform required for amylopectin synthesis in Chlamydomonas reinhardtii.” Frontiers in plant science vol. 14 1201386. 31 May. 2023, doi:10.3389/fpls.2023.1201386
 Figure
1.1 The pattern diagram of the DBTL cycle
Figure
1.1 The pattern diagram of the DBTL cycle 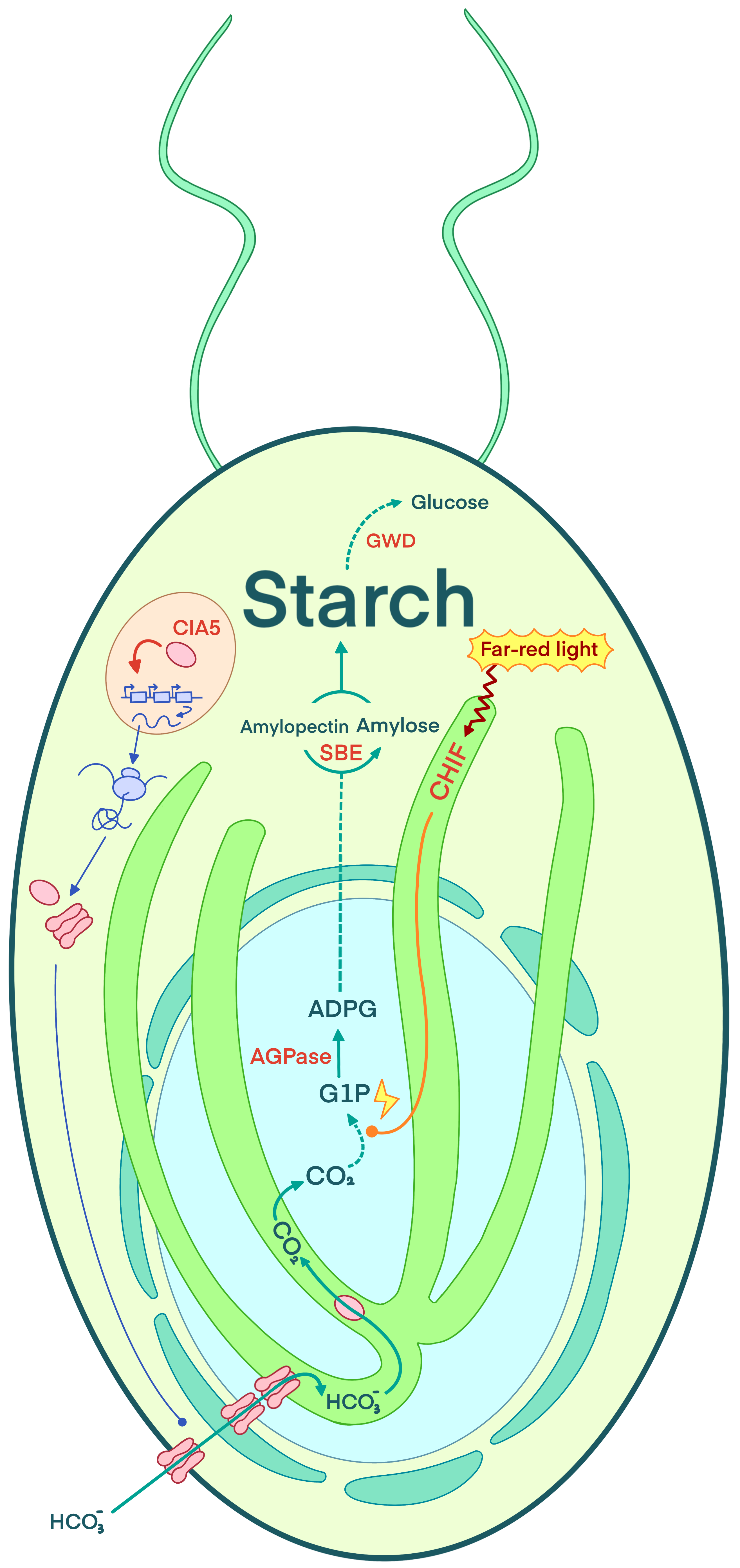

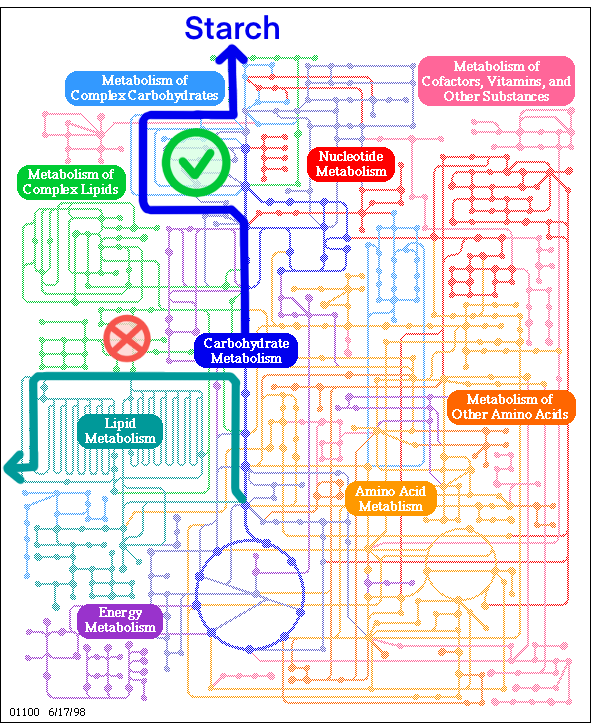 Figure2.1
Click Here to view the page[1]
Figure2.1
Click Here to view the page[1]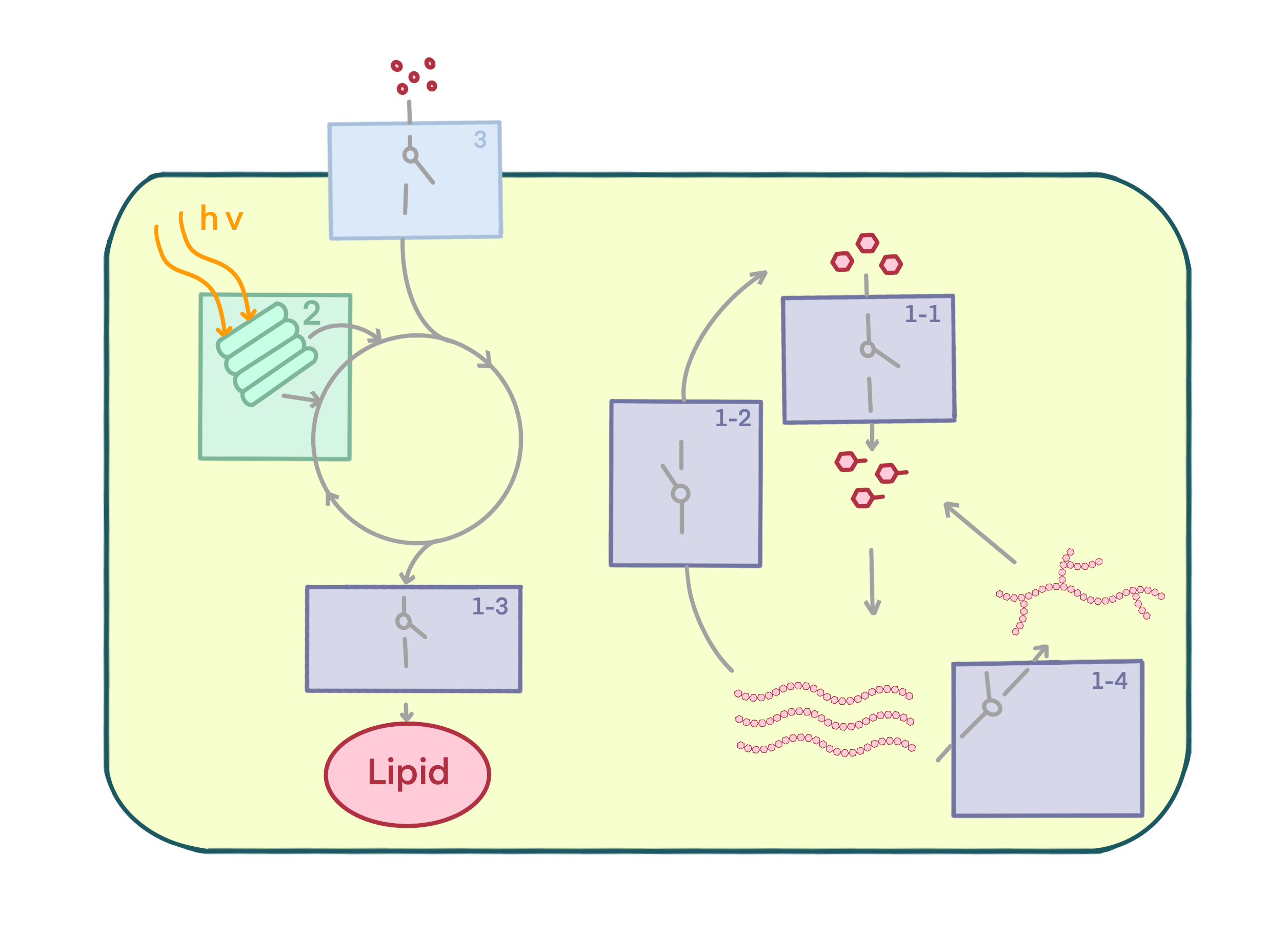 Figure
2.2 Schematic diagram of regulatory modules and metabolic circuit
Figure
2.2 Schematic diagram of regulatory modules and metabolic circuit
 Figure 2.4 AGPase
and gwd activity’s influence on starch yield
Figure 2.4 AGPase
and gwd activity’s influence on starch yield Figure 2.5 Effect of
branching enzyme activity on starch branching
Figure 2.5 Effect of
branching enzyme activity on starch branching 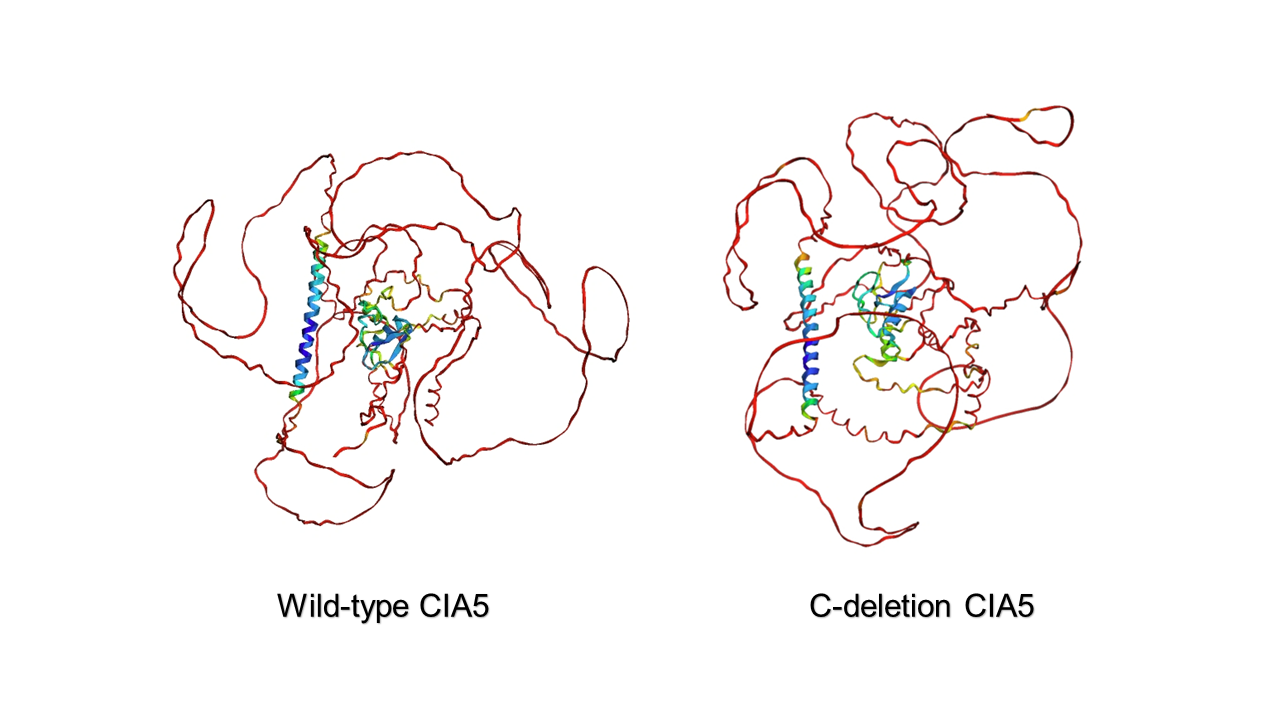 Figure
2.6 Prediction of the structure of wild-type and C-deletion CIA5 protein
Figure
2.6 Prediction of the structure of wild-type and C-deletion CIA5 protein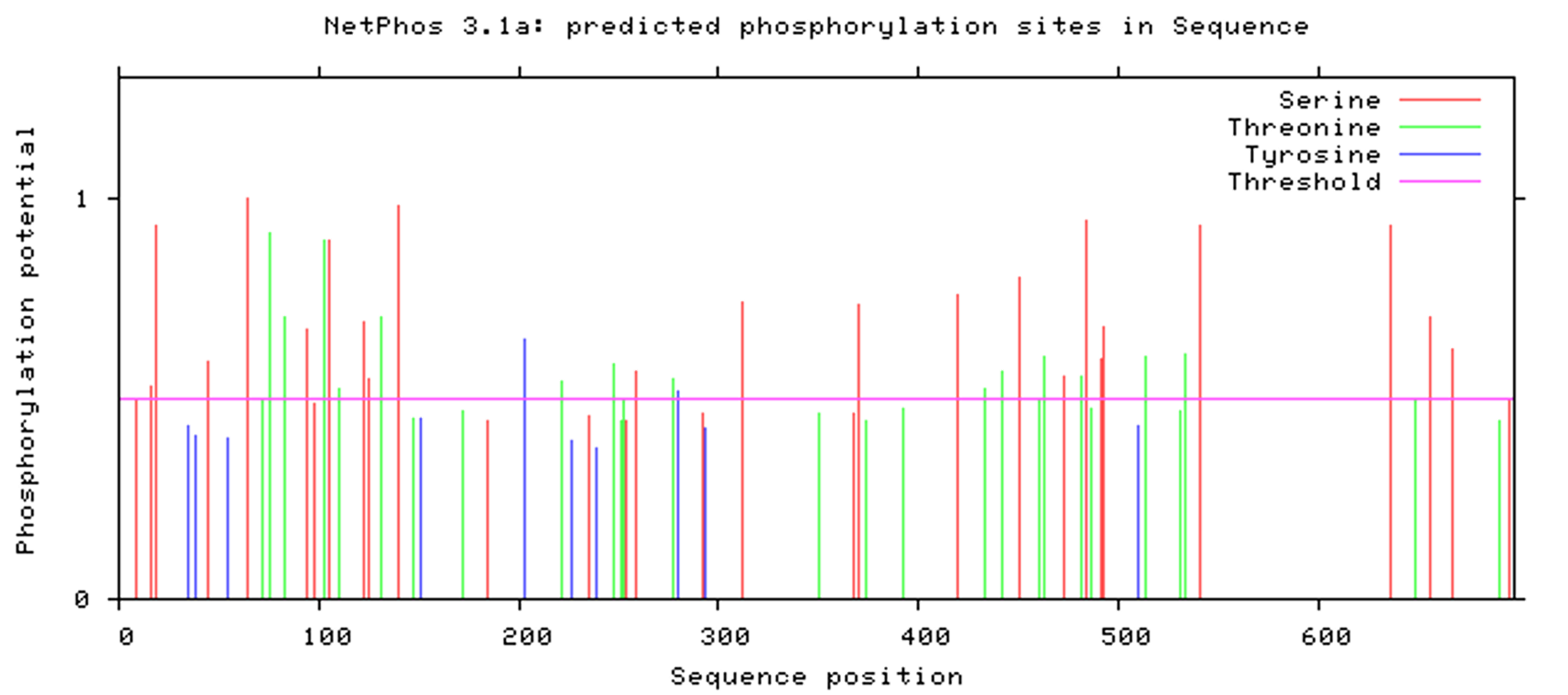 Figure 2.7 Prediction of the phosphorylation site at the C-terminus of the CIA5 protein
Figure 2.7 Prediction of the phosphorylation site at the C-terminus of the CIA5 protein 
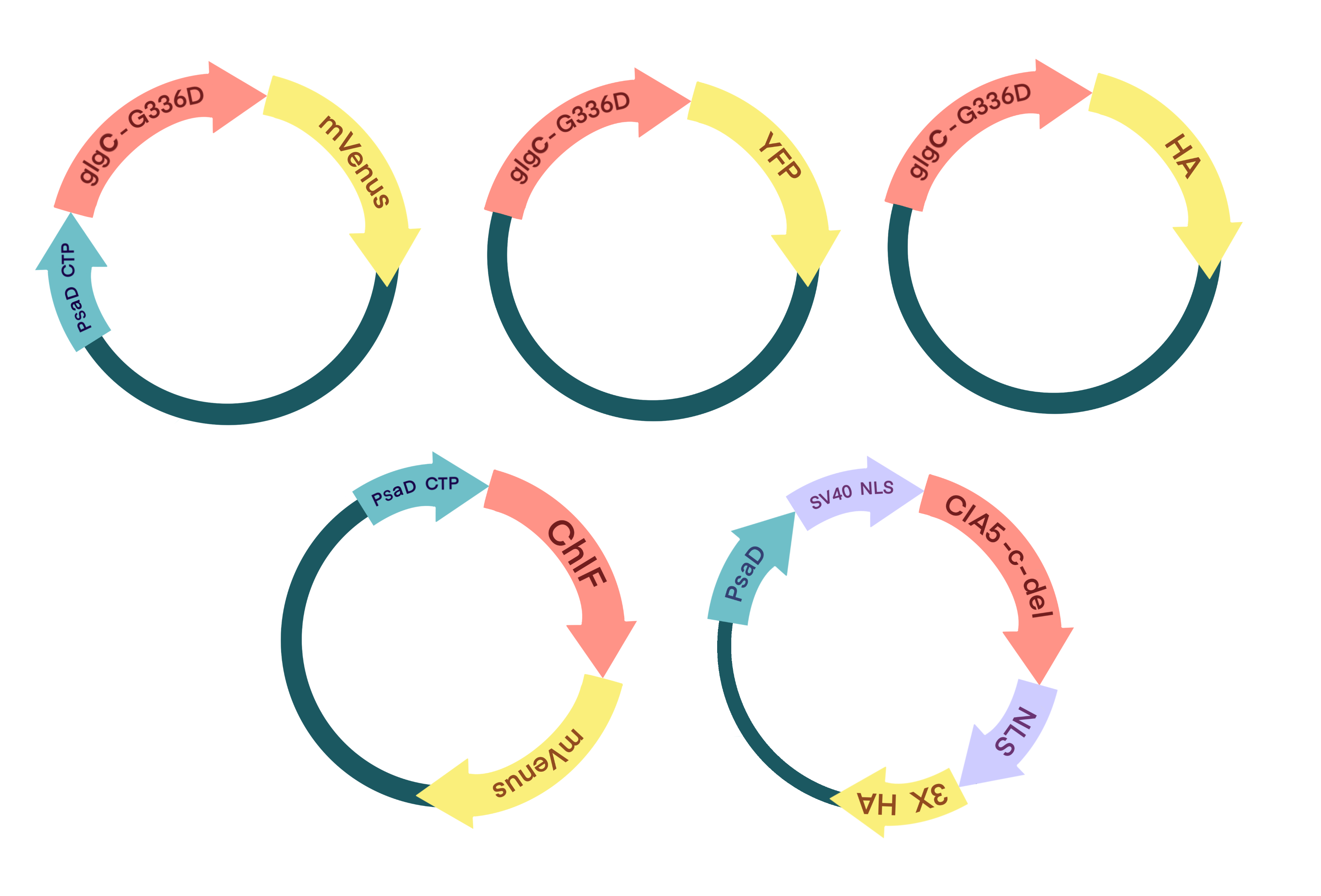 All of
plasmid profiles in three modules. (Promoters, resistance elements, etc., not drawn)
All of
plasmid profiles in three modules. (Promoters, resistance elements, etc., not drawn)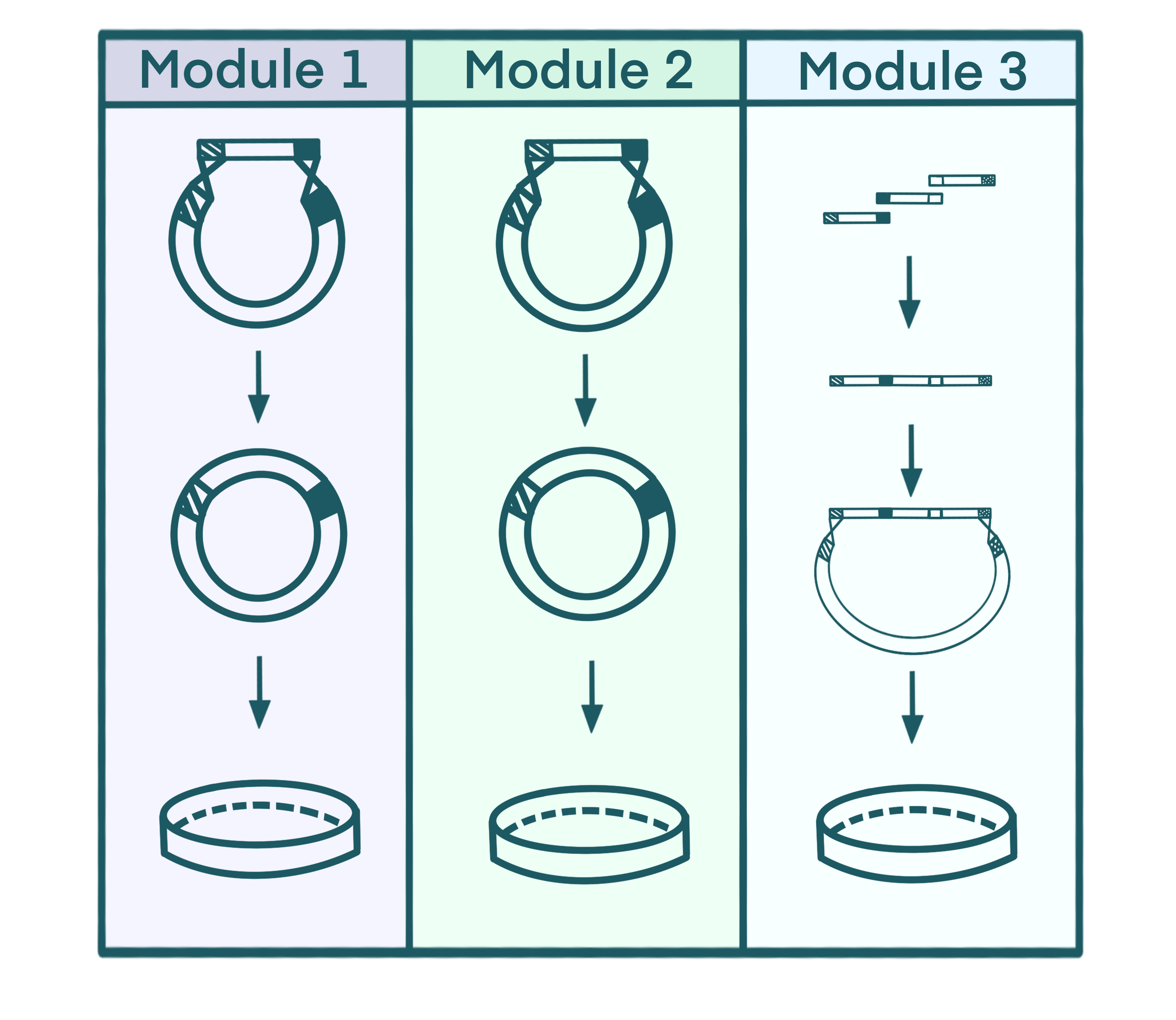 Figure
3.1 Gene assembly paradigm of Module1-3
Figure
3.1 Gene assembly paradigm of Module1-3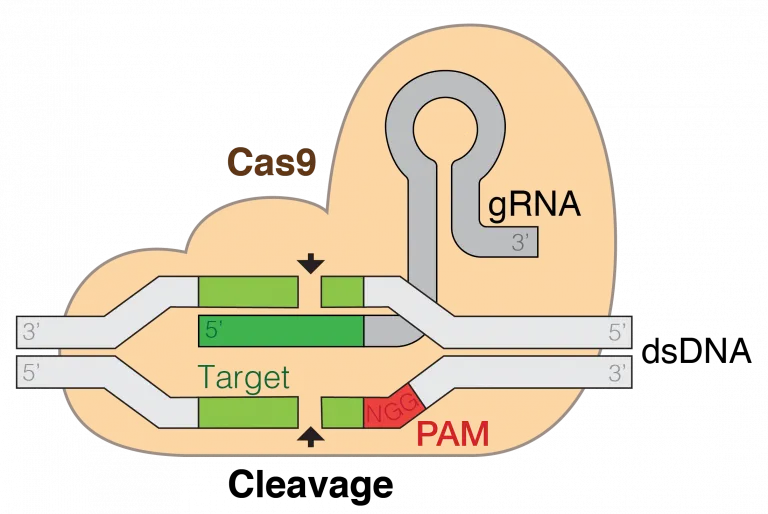 Figure
3.2 Paradigm of
CRISPR/Cas9[5]
Figure
3.2 Paradigm of
CRISPR/Cas9[5]
 Figure
3.3 gel-shift conformation of corresponding mutants.
Figure
3.3 gel-shift conformation of corresponding mutants.








 Figure
3.8 Percent of Amylose to total starch
Figure
3.8 Percent of Amylose to total starch  Figure 3.9 Phenotypic characterization of starches produced by mutant
Figure 3.9 Phenotypic characterization of starches produced by mutant  Figure 3.10 data for
measuring the OD values of Chlamydomonas
Figure 3.10 data for
measuring the OD values of Chlamydomonas Figure
3.11 fluorescence intensity check on YFP
Figure
3.11 fluorescence intensity check on YFP Figure 3.12 Western blot result on Chif-YFP fused protein
Figure 3.12 Western blot result on Chif-YFP fused protein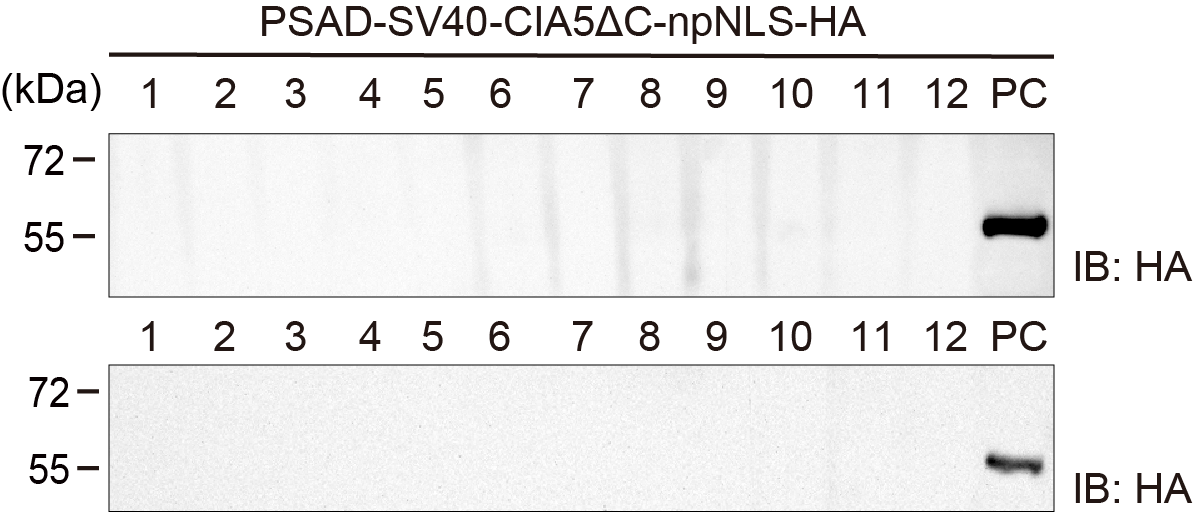 Figure
3.13 western blot screening for CIA5ΔC protein expression
Figure
3.13 western blot screening for CIA5ΔC protein expression Figure
3.14 Learning-cycle
Figure
3.14 Learning-cycle